Research Articles
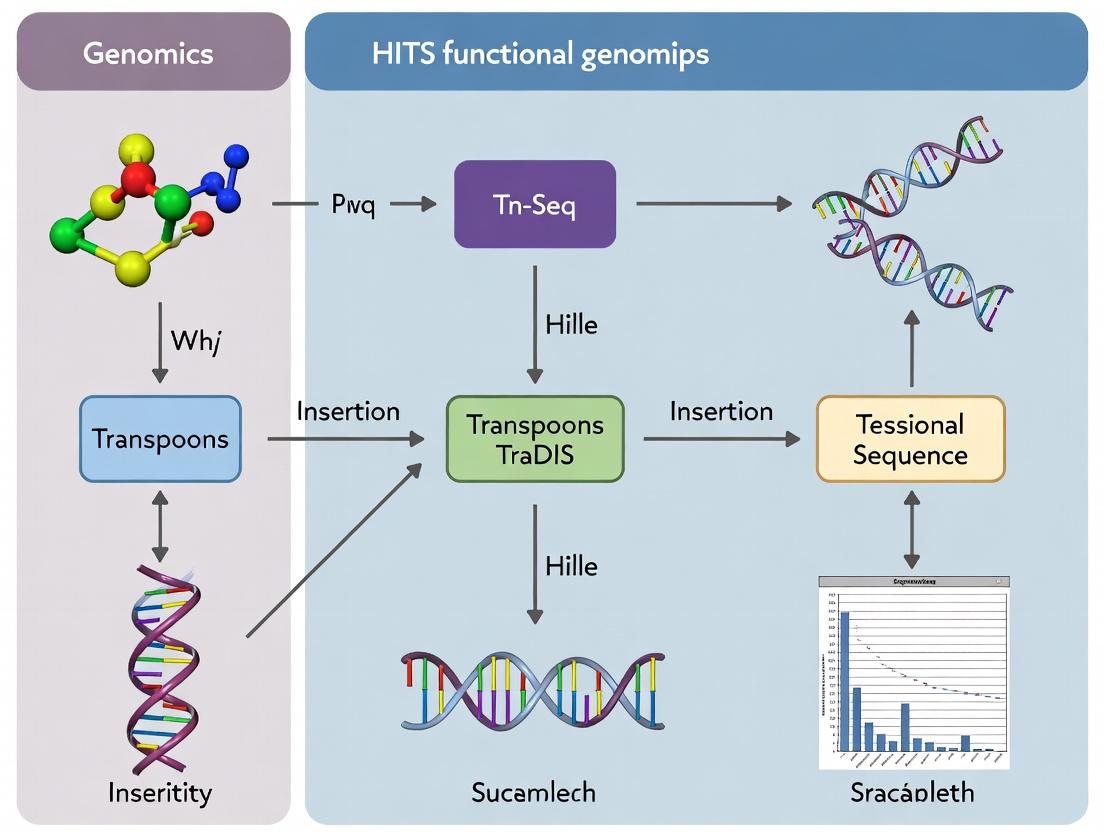
Tn-Seq vs. TraDIS vs. HITS: A Comparative Guide to High-Throughput Functional Genomics Methods
This comprehensive guide explores Tn-Seq, TraDIS, and HITS, three cornerstone techniques in high-throughput functional genomics.
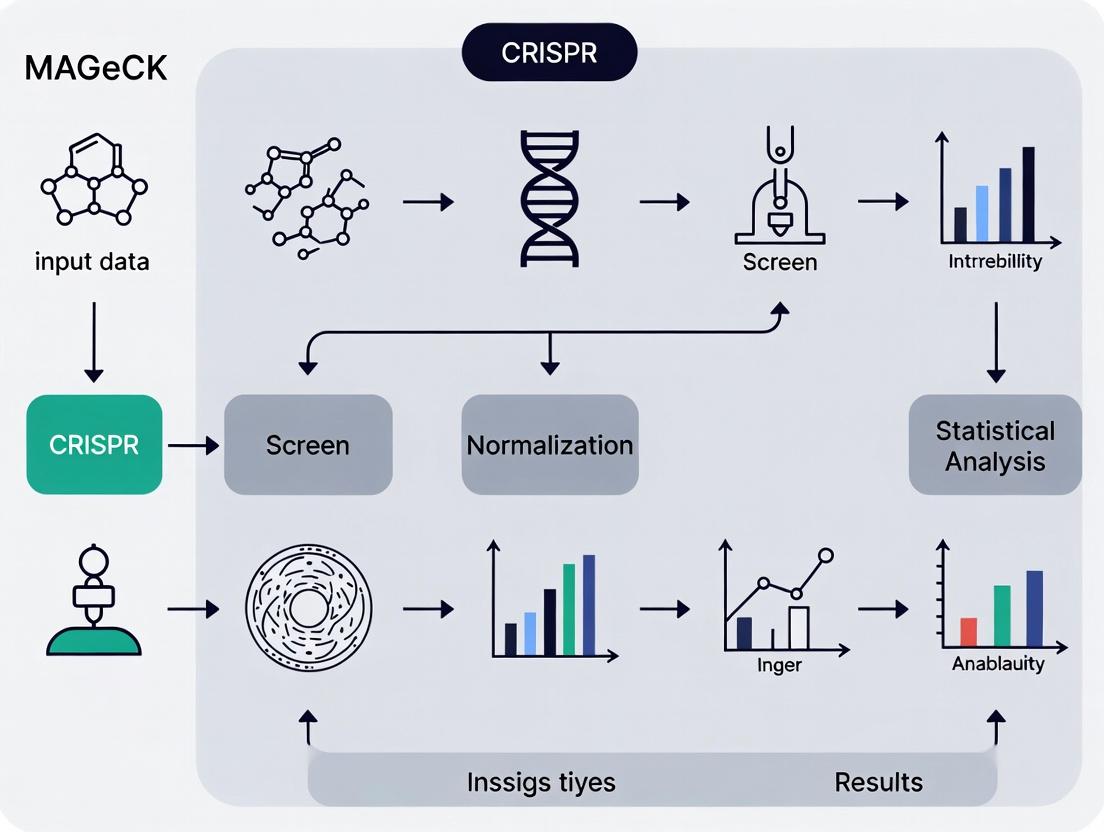
Mastering CRISPR Screen Analysis: A Complete MAGeCK Tutorial from Raw Data to Biological Insights
This comprehensive guide provides researchers, scientists, and drug development professionals with a complete workflow for analyzing CRISPR screening data using MAGeCK.
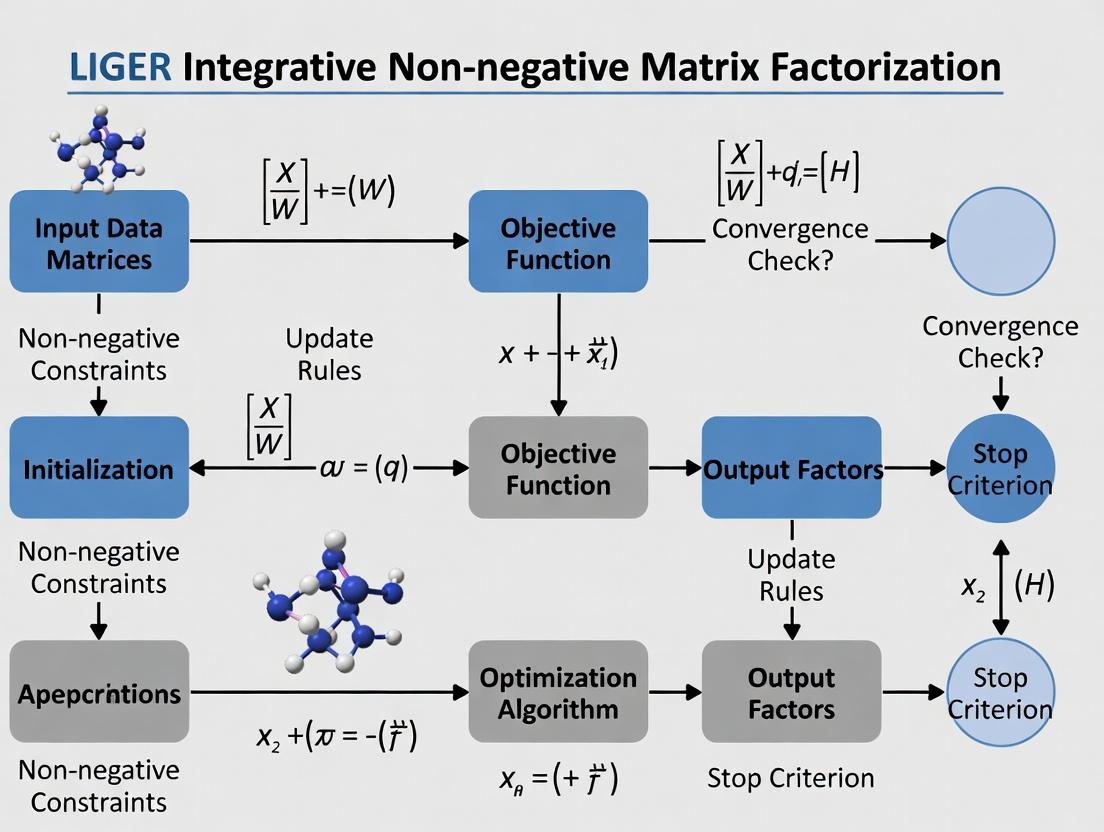
LIGER Integrative Non-Negative Matrix Factorization: A Complete Guide for Multi-Omics Data Analysis in Biomedical Research
This article provides a comprehensive exploration of LIGER (Linked Inference of Genomic Experimental Relationships), a powerful integrative non-negative matrix factorization (iNMF) framework.
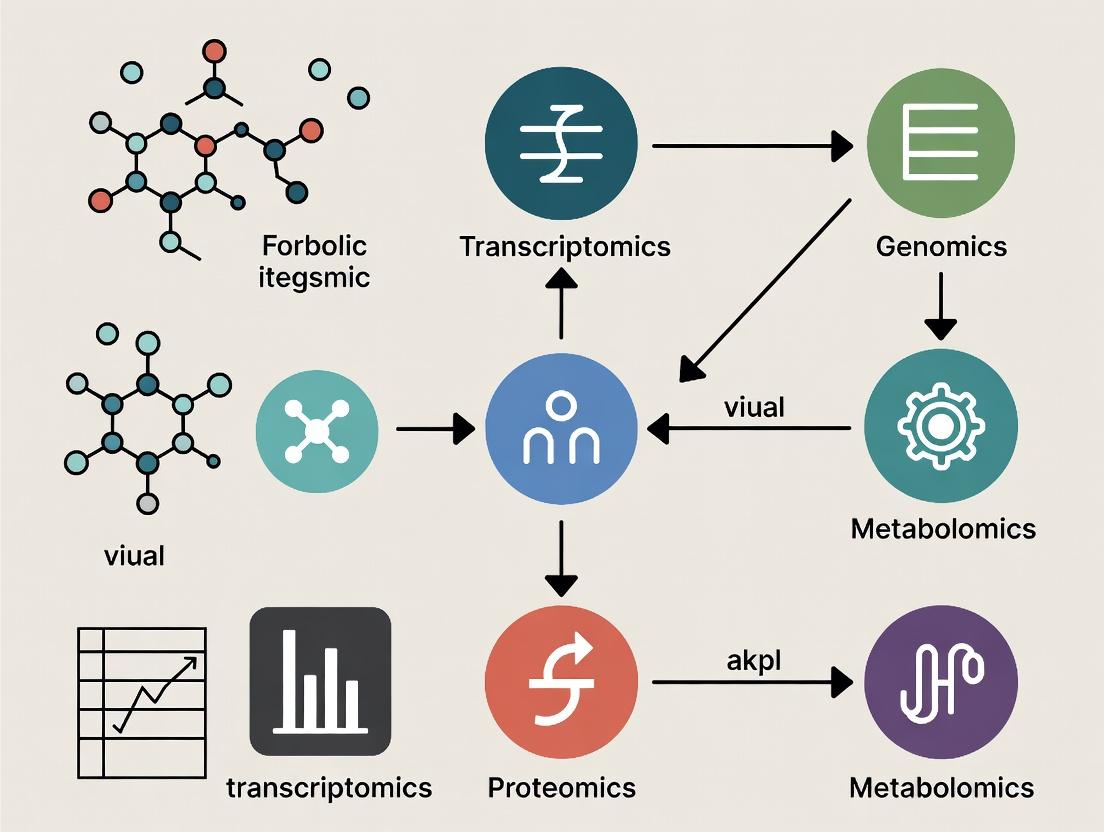
Multi-Omics Integration: A Comprehensive Guide to Methods, Applications, and Best Practices for Biomedical Research
This article provides a comprehensive overview of multi-omics integration methods, tailored for researchers, scientists, and drug development professionals.
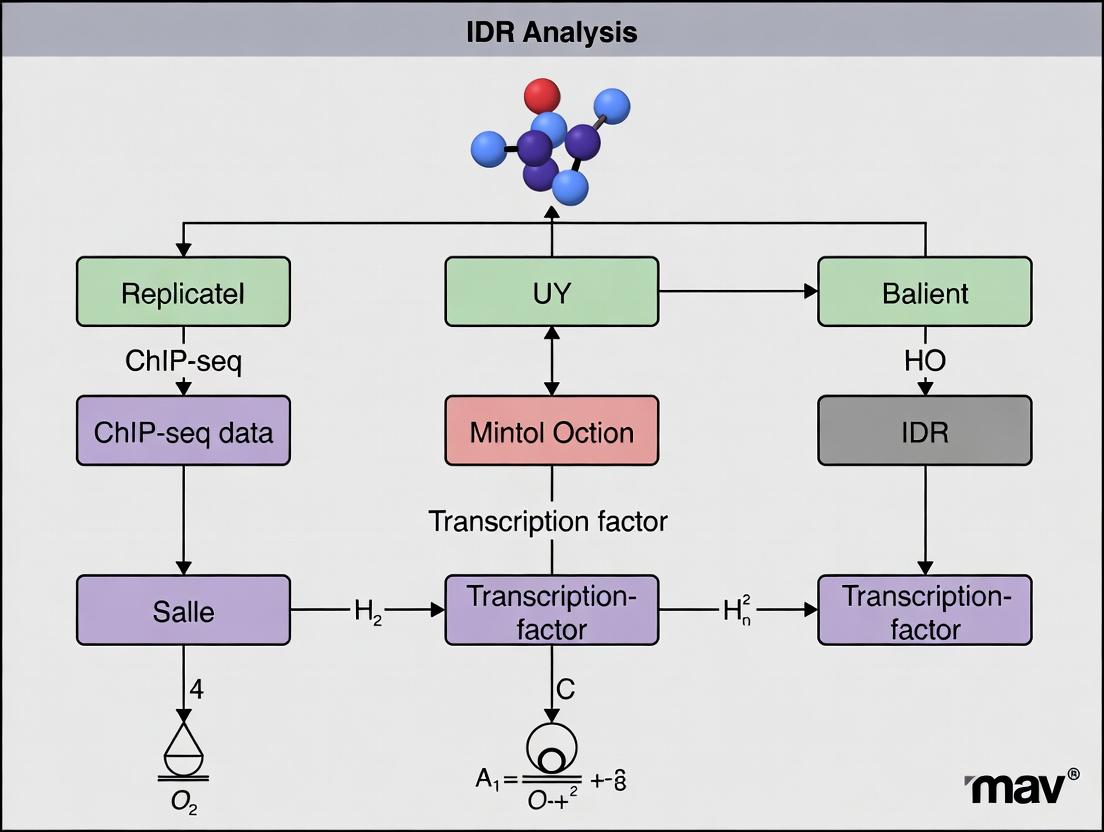
Mastering IDR Analysis: The Complete Guide to Reproducible Transcription Factor ChIP-seq Peak Calling
This comprehensive guide provides researchers and drug development scientists with a complete framework for performing Irreproducible Discovery Rate (IDR) analysis on replicated transcription factor (TF) ChIP-seq experiments.
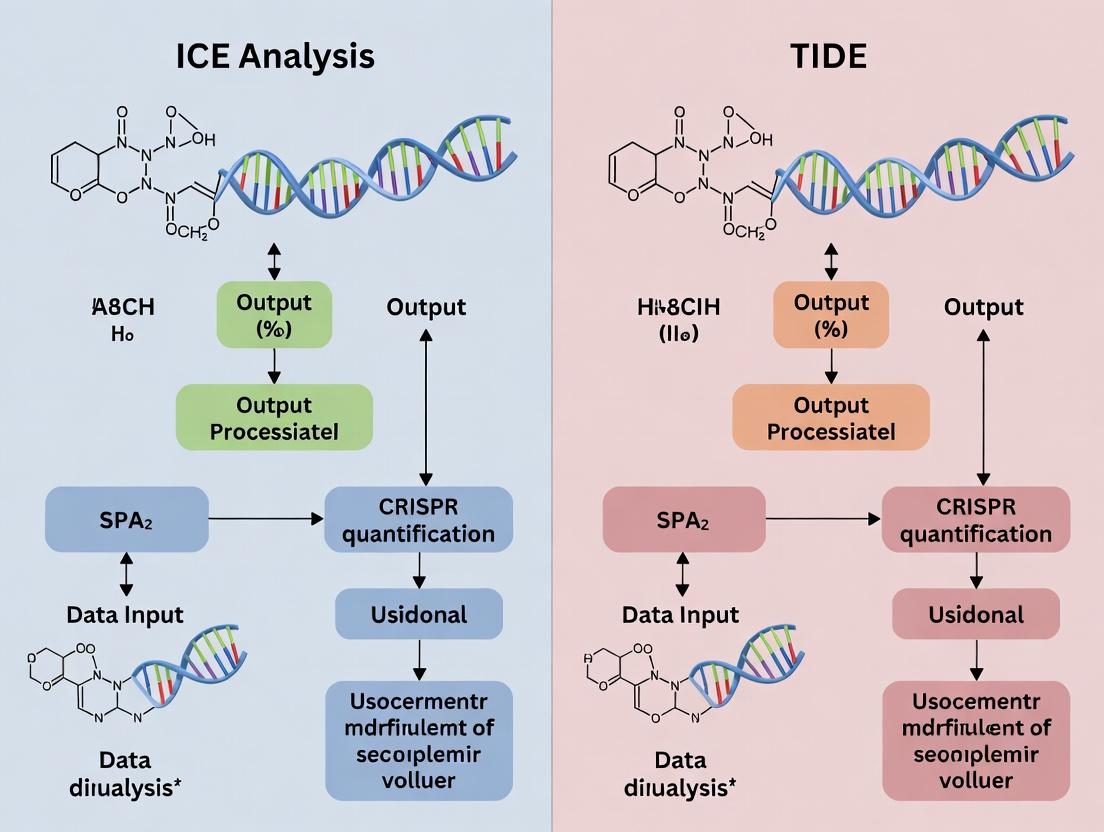
ICE vs TIDE for CRISPR Analysis: A Comprehensive Guide for Accurate Edit Quantification
This article provides a detailed comparison of the two predominant methods for analyzing CRISPR-Cas9 editing efficiency: Inference of CRISPR Edits (ICE) and Tracking of Indels by Decomposition (TIDE).
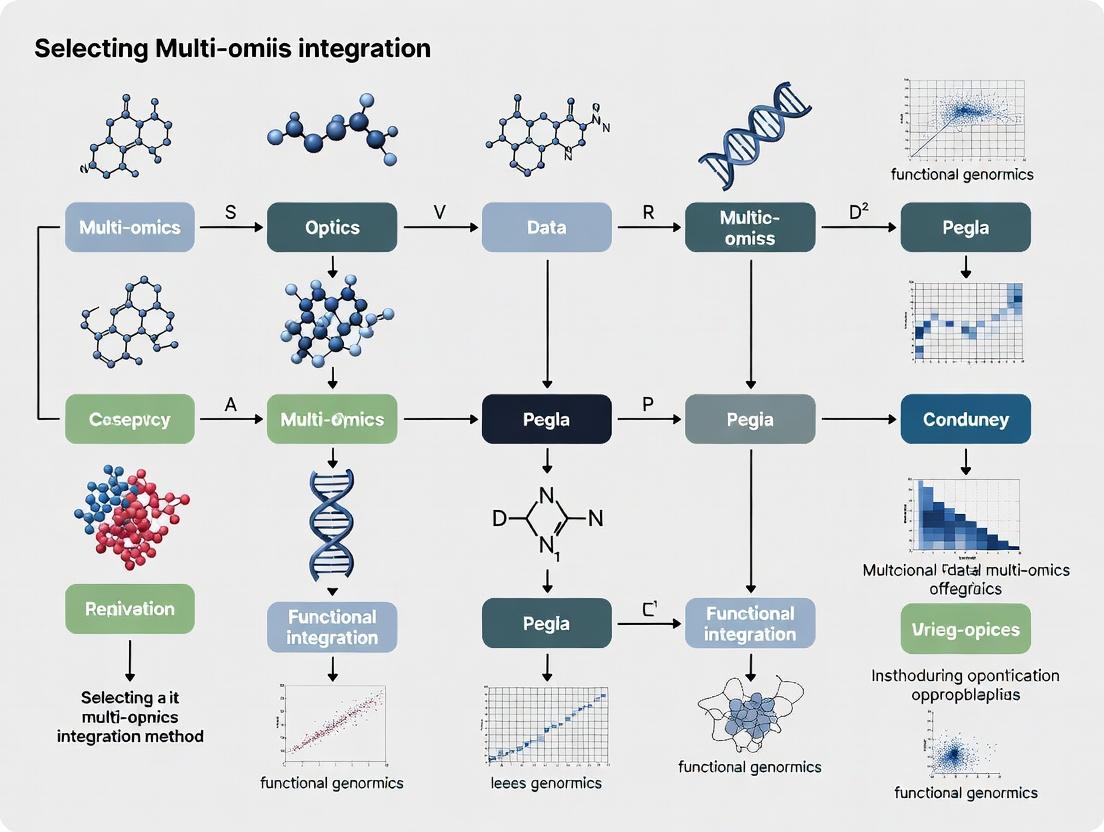
Multi-Omics Integration Demystified: A Practical Guide to Choosing the Right Method for Your Research
This comprehensive guide empowers researchers, scientists, and drug development professionals to navigate the complex landscape of multi-omics data integration.
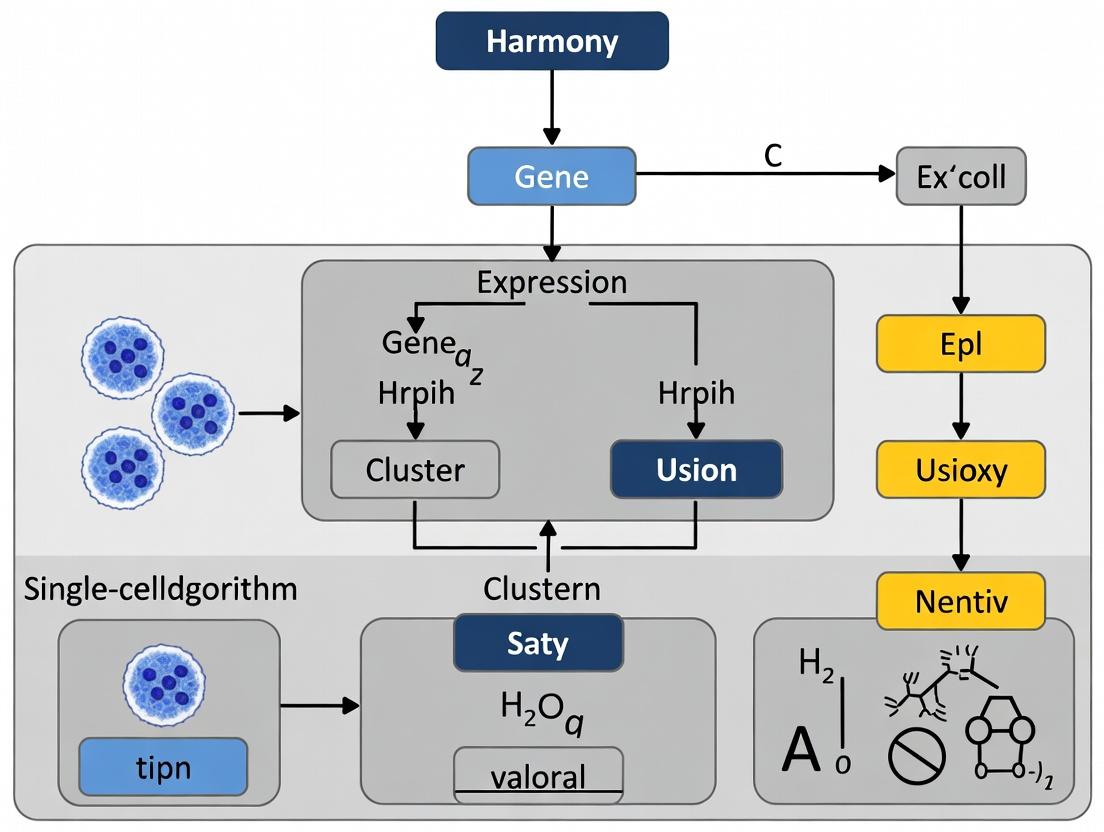
Harmony Algorithm: A Comprehensive Guide to Single-Cell Data Integration for Biomedical Research
This comprehensive guide explores the Harmony algorithm for single-cell RNA sequencing data integration, providing researchers and drug development professionals with foundational understanding, practical application workflows, troubleshooting strategies, and comparative validation...
Mastering Harman Batch Effect Correction: A Complete Guide to Constraint Settings for Robust Omics Data Analysis
This comprehensive guide explores the critical role of constraint settings in Harman batch effect correction for genomic, transcriptomic, and proteomic data.
Missing in Multi-Omics: A Comprehensive Guide to Handling Missing Values in Genomic, Transcriptomic, and Proteomic Data
Missing data is an omnipresent challenge in multi-omics studies, threatening the validity of integrative analysis and downstream biological discovery.